Two members of the University of Rochester’s Department of Biology have received expedited funding awards from the National Science Foundation (NSF) to study biological processes involved in COVID-19.
As labs reopen across the University, Dragony Fu, an assistant professor of biology, and Jack Werren, the Nathaniel and Helen Wisch Professor of Biology, will apply their expertise in cellular and evolutionary biology to research proteins involved in infections from COVID-19, which is caused by the novel coronavirus SARS-CoV-2. The funding is part of the NSF’s Rapid Response Research (RAPID) program to mobilize funding for high priority projects.
“At this point, combating this pandemic is an ‘all-hands-on-deck’ situation,” says Elaine Sia, professor and chair of biology. “Researchers in the biology department at the University, like biologists everywhere, have been learning all we can about the SARS-CoV2 virus.”
Rochester Restart
Stay up to date on the University’s plans to safely phase in campus operations.
Coronavirus concern
If you’re concerned about sympotoms of potential exposure to coronavirus, visit the University’s COVID-19 website for guidance and information.
By better understanding the specific biological mechanisms and proteins involved in COVID-19 infection, scientists will better be able to develop effective treatments and vaccines to fight the disease.
Connections between coronavirus, proteins, and RNA
The COVID-19 virus makes proteins called proteases that act like scissors to cut the virus’s own proteins into smaller, functional pieces that are required for viral infection. A research group at the University of California San Francisco recently discovered that the main protease of SARS-CoV2 interacts with a human protein called TRNA Methyltransferase 1 (TRMT1), which Dragony Fu’s lab had been studying well before the COVID-19 pandemic.
“The central goal of our lab is to understand the functions of proteins that modify RNA,” Fu says. “TRMT1 happens to be one of the main RNA modification proteins we study, so it was quite serendipitous that it is connected to the COVID-19 virus because we have already established tests to measure TRMT1 function in human cells.”
With his grant, Fu is partnering with researchers at the French National Centre for Scientific Research to study why SARS-CoV2 interacts with TRMT1 and how the interaction affects both the virus and human cells. Fu hypothesizes that the coronavirus protease cuts TRMT1, preventing the protein from modifying RNA as it normally does, and, in turn, compromising RNA’s function in protein synthesis.
“Dr. Fu is well positioned to carry out these studies using the rigorous biochemical and molecular approaches that have characterized his previous research endeavors,” Sia says. “In addition, he has the admirable ability to establish effective collaborations across the world, which has allowed him to tackle some really difficult problems in the past.”
Fu and his collaborators will use molecular biology techniques to deliver the viral protease gene into human lung cells. They will then use antibodies to detect TRMT1 to determine whether the protein is being cut by the viral protease. If so, they will use biochemical assays to analyze how the viral protease affects TRMT1 function, RNA modification, and protein synthesis.
“We will be able to use this knowledge to discover new connections between the virus and the host human cell,” Fu says. “These novel links could serve as potential therapeutic targets for treatment of COVID-19 infection.”
Read more about Fu’s project.
Using a past approach to explore proteins and enzyme receptors
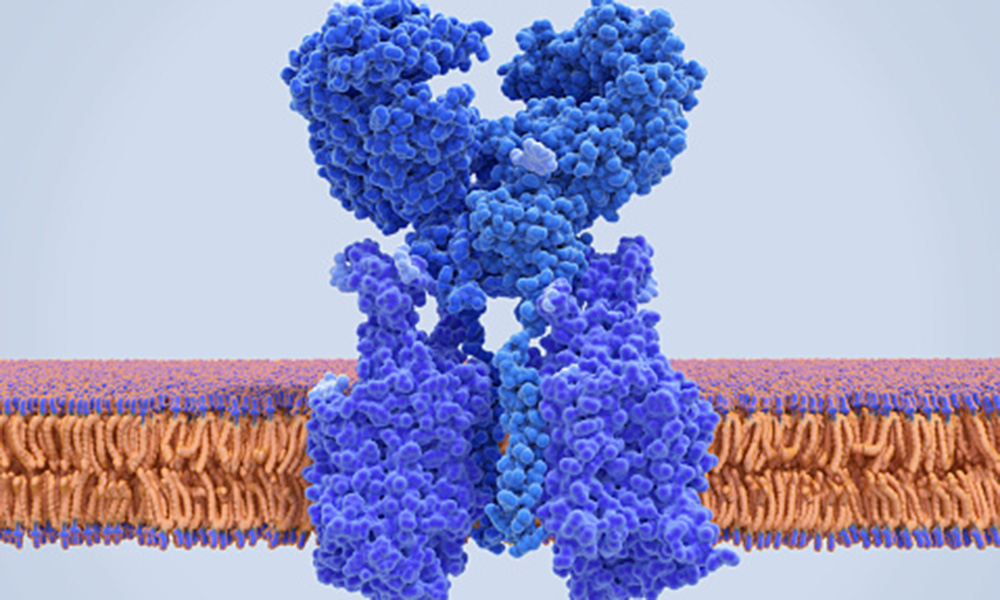
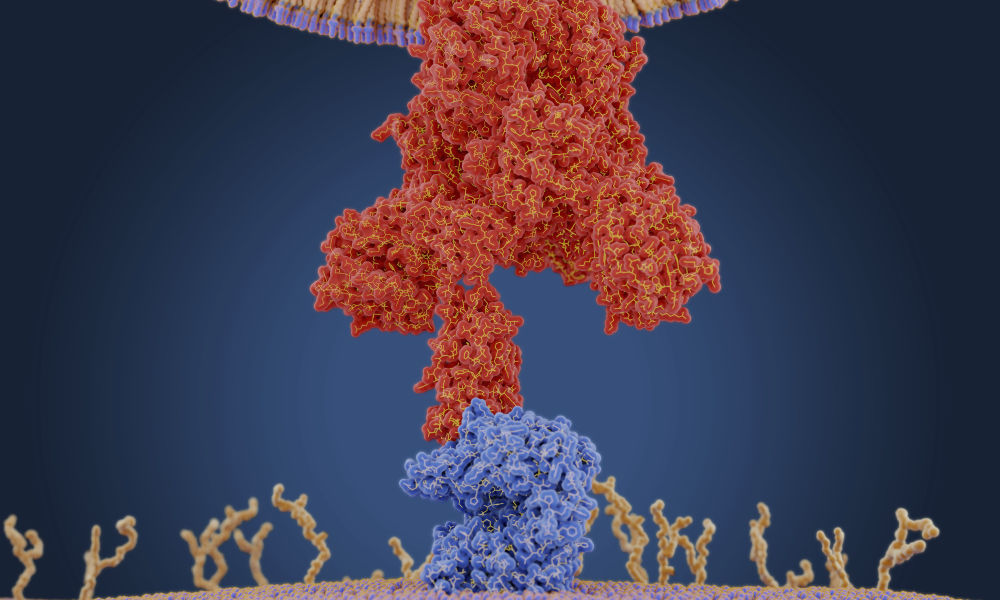
Once the COVID-19 virus enters the body, it binds to cells via an enzyme receptor called Angiotensin-converting enzyme 2 (ACE2), which is located on cell surfaces.
Jack Werren is using his grant to identify potential interactions between ACE2 and other human proteins that are involved in human health problems associated with COVID-19 infection.
“Dr. Werren is an evolutionary geneticist who has applied his considerable skills to a variety of important questions over the course of his career, many of them involving the interaction of genomes in symbiotic or parasitic relationships,” Sia says. “An examination of the interaction between the SARS-CoV2 virus and the human host cell was, therefore, a natural fit for his expertise.”
Werren is using a novel computational and evolutionary approach called “evolutionary rate correlation” (ERC) to identify proteins that may coevolve and interact with the ACE2 receptor on human cells. Proteins that interact with each other are likely to show similar rates of change during evolution. For example, if you look at a phylogenetic tree depicting the evolutionary relationships among mammals, when one protein evolves slowly in a particular species, a protein with which it interacts will tend to evolve slowly as well and vice versa.
Werren has previously applied the ERC method to proteins that interact to produce mitochondria—the cellular structure that, among other functions, produces energy for cells. He found that the ERC method was very accurate in predicting proteins known to interact with mitochondria.
When COVID-19 hit, Werren says, “we decided to see if the ERC approach could be useful in identifying protein interactions relevant to the severe symptoms of the disease.”
Working with recent Rochester graduate Austin Varela ’20, who joined the ERC project when he was a sophomore, Werren first focused on identifying proteins that coevolve—and therefore might interact—with ACE2. He has detected several proteins not previously known to interact with ACE2, but that show strong signals of coevolving with ACE2. Several of the proteins are potentially relevant to afflictions that people infected with COVID-19 experience, such as blood clotting and arterial disease.
“Our goal is to characterize these interactions further and quickly publish the findings, in case the information will be useful in identifying targets for therapeutic intervention,” Werren says.
Read more about Werren’s project.
Read more
 COVID-19: What’s RNA research got to do with it?
COVID-19: What’s RNA research got to do with it?Rochester research into RNA structure and function provides key information for developing coronavirus treatments.

Evolution’s New Era
Rochester biologists are leaders in a paradigm shift as scientists explore an evolutionary arms race that’s not between organisms but within them.

Researchers adapt to precautions
Time sharing, staggered shifts, and reconfigured spaces are among the adaptations the University has made for research to resume.